Discovering and documenting
the diversity of marine life
A new program at the Natural History Museum of Los Angeles County aims to develop research and monitoring tools to enormously accelerate our ability to discover and investigate marine invertebrate biodiversity of California’s nearshore waters. The Diversity Initiative for the Southern California Ocean (DISCO) is based in the NHM’s Marine Biodiversity Center, and incorporates partnerships with organizations both inside and outside of the Museum.
There are good things to see in the tidepools and there are exciting and interesting thoughts to be generated from the seeing. Every new eye applied to the peep hole which looks out at the world may fish in some new beauty and some new pattern, and the world of the human mind must be enriched by such fishing.
John Steinbeck. 1948.
Foreword to:
E.F. Ricketts & J. Calvin, Between Pacific Tides.
Why a new diversity intiative?
We know that our oceans are changing quickly, and our coastal biota are responding. Successful management requires that we know what species are where, and how their distributions change with time.
Current monitoring technology, based on specialist identifications of individual organisms, just cannot keep pace. Standard approaches to ecological research and monitoring efforts that look at marine species distributions depend on highly trained taxonomists to sort and identify the animals in environmental samples. Identification of marine macroinvertebrates is challenging because of their small size, complex morphology, and extreme diversity. Therefore, because of the expertise and time required, relatively few sites can be sampled, and often studies are restricted to a handful of “indicator” species, rather than comprehensively looking at the community. This cripples our ability to see the widespread but subtle changes to our coastal biota that accompany variations in ocean temperature and acidity. We need quicker and more effective ways to identify the species in marine samples.
New genetic approaches
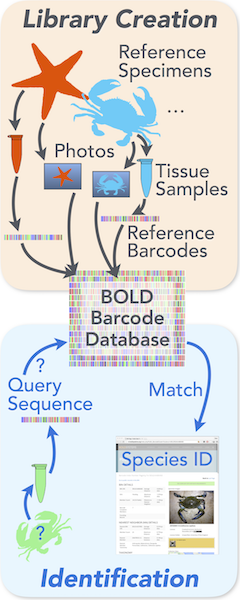
Over the last decade, molecular identification tools, called “genetic barcoding,” have been developed to identify the species of any specimen based on its DNA. Species share a handful of particular genes that exhibit just the right amount of variation for dependable identification. Individual specimens can be identified to species by sequencing a single (or very few) short gene sequences and then comparing those sequences against a reference library of sequences from independently (non-genetically) identified specimens.
Of course, for that system to work, the reference library of “barcode” sequences (the Barcode of Life Database [BOLD] run by the University of Guelph) must contain at least one reference sequence for each species of interest. Expert taxonomists must reputably identify specimens, get the appropriate DNA sequences from them, deposit the sequences in BOLD, and deposit the voucher specimens in a museum for permanent storage.
Building the library
Sadly, the estimated 3,000 to 5,000 marine invertebrate species of Southern California are poorly represented in the BOLD reference library. NHM’s DISCO project aims to fill in the missing entries. We will use a combination of existing NHM specimens, new collections, and external contributions. NHM taxonomists have the expertise and professional collaborators necessary to undertake and complete this task. All specimens collected and processed in this project will be archived as permanent voucher specimens at the NHM and all sequences will be publicly available at BOLD.
Dramatic inventory improvements
Being able to attribute an individual specimen to a species based on its genetic sequence can be useful. But much more exciting are the prospects offered by combining a well-populated barcode reference library with modern high-throughput genetic sequencing.
It is now possible to extract DNA from an entire community at once (e.g. combined scrapings from a floating dock) and then retrieve all of the unique sequences in the entire community with a single run of a high-throughput sequencer. Even more extraordinary, just sampling DNA from the seawater around a community can yield genetic diversity information. Sequencing of DNA from the environment, or eDNA, is possible because all organisms naturally shed DNA into the environment around them. There can be enough DNA in seawater samples that it is unnecessary to sample the organisms directly.
These approaches yield a list of unique genetic sequences. By searching those sequences against a well-populated reference library, the list of sequences can be interpreted as a list of species.
Whole-community and eDNA technology is at the prototype stage for coastal marine systems. Another goal of NHM’s DISCO project is to work with partner organizations to develop this technology to the point that it is routinely applicable in marine research and monitoring.
Being able to inventory full species diversity quickly and efficiently will permit far more frequent sampling in monitoring applications, and much more taxonomically broad coverage in research programs.
Who we are
The DISCO program comes from the marine invertebrate group at the Natural History Museum of Los Angeles County.
DISCO is led by Regina Wetzer (Associate Curator) and based out of the Marine Biodiversity Center (MBC). Other key personnel include Dean Pentcheff (Project Coordinator), Leslie Harris (Collections Manager, Polychaetes), Kathy Omura (Collections Manager, MBC), Adam Wall (Collections Manager, Crustacea), and Jenessa Wall (Assistant Collections Manager, MBC).
Further information
Additional information about genetic barcoding and its applications to ecological research and monitoring (both marine and terrestrial) can be found in the following references.
Blaxter, M. 2016. Imagining Sisyphus happy: DNA barcoding and the unnamed majority. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150329.
Bohmann, K., A. Evans, M. T. P. Gilbert, G. R. Carvalho, S. Creer, M. Knapp, D. W. Yu, and M. de Bruyn. 2014. Environmental DNA for wildlife biology and biodiversity monitoring. Trends in Ecology & Evolution 29:358–367.
Cristescu, M. E. 2014. From barcoding single individuals to metabarcoding biological communities: towards an integrative approach to the study of global biodiversity. Trends in Ecology & Evolution 29:566–571.
Gibson, J., S. Shokralla, T. M. Porter, I. King, S. van Konynenburg, D. H. Janzen, W. Hallwachs, and M. Hajibabaei. 2014. Simultaneous assessment of the macrobiome and microbiome in a bulk sample of tropical arthropods through DNA metasystematics. Proceedings of the National Academy of Sciences 111:8007–8012.
Goldberg, C. S., K. M. Strickler, and D. S. Pilliod. 2015. Moving environmental DNA methods from concept to practice for monitoring aquatic macroorganisms. Biological Conservation 183:1–3.
Hajibabaei, M. 2012. The golden age of DNA metasystematics. Trends in Genetics 28:535–537.
Hajibabaei, M., D. J. Baird, N. A. Fahner, R. Beiko, and G. B. Golding. 2016. A new way to contemplate Darwin’s tangled bank: how DNA barcodes are reconnecting biodiversity science and biomonitoring. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150330.
Hebert, P. D. N., P. M. Hollingsworth, and M. Hajibabaei. 2016a. From writing to reading the encyclopedia of life. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150321.
Hebert, P. D. N., S. Ratnasingham, E. V. Zakharov, A. C. Telfer, V. Levesque-Beaudin, M. A. Milton, S. Pedersen, P. Jannetta, and J. R. deWaard. 2016b. Counting animal species with DNA barcodes: Canadian insects. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150333.
Ji, Y., L. Ashton, S. M. Pedley, D. P. Edwards, Y. Tang, A. Nakamura, R. Kitching, P. M. Dolman, P. Woodcock, F. A. Edwards, T. H. Larsen, W. W. Hsu, S. Benedick, K. C. Hamer, D. S. Wilcove, C. Bruce, X. Wang, T. Levi, M. Lott, B. C. Emerson, and D. W. Yu. 2013. Reliable, verifiable and efficient monitoring of biodiversity via metabarcoding. Ecology Letters 16:1245–1257.
Kelly, R. P., J. L. O’Donnell, N. C. Lowell, A. O. Shelton, J. F. Samhouri, S. M. Hennessey, B. E. Feist, and G. D. Williams. 2016. Genetic signatures of ecological diversity along an urbanization gradient. PeerJ 4:e2444.
Kelly, R. P., J. A. Port, K. M. Yamahara, and L. B. Crowder. 2014. Using environmental DNA to census marine fishes in a large mesocosm. PLoS ONE 9:e86175.
La Salle, J., K. J. Williams, and C. Moritz. 2016. Biodiversity analysis in the digital era. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150337.
Leray, M., and N. Knowlton. 2015. DNA barcoding and metabarcoding of standardized samples reveal patterns of marine benthic diversity. Proceedings of the National Academy of Sciences 112:2076–2081.
Leray, M., and N. Knowlton. 2016. Censusing marine eukaryotic diversity in the twenty-first century. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150331.
Miller, S. E., A. Hausmann, W. Hallwachs, and D. H. Janzen. 2016. Advancing taxonomy and bioinventories with DNA barcodes. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150339.
Pansu, J., S. De Danieli, J. Puissant, J.-M. Gonzalez, L. Gielly, T. Cordonnier, L. Zinger, J.-J. Brun, P. Choler, P. Taberlet, and L. Cécillon. 2015. Landscape-scale distribution patterns of earthworms inferred from soil DNA. Soil Biology and Biochemistry 83:100–105.
Ratnasingham, S., and P. D. N. Hebert. 2007. BOLD: The Barcode of Life Data System (http://www. barcodinglife.org). Molecular Ecology Notes 7:355–364.
Ratnasingham, S., and P. D. N. Hebert. 2013. A DNA-Based Registry for All Animal Species: The Barcode Index Number (BIN) System. PLoS ONE 8:e66213.
Shokralla, S., T. M. Porter, J. F. Gibson, R. Dobosz, D. H. Janzen, W. Hallwachs, G. B. Golding, and M. Hajibabaei. 2015. Massively parallel multiplex DNA sequencing for specimen identification using an Illumina MiSeq platform. Scientific Reports 5:9687.
Shokralla, S., G. Singer, and M. Hajibabaei. 2010. Direct PCR amplification and sequencing of specimens’ DNA from preservative ethanol. BioTechniques 48:233–234.
Shokralla, S., J. L. Spall, J. F. Gibson, and M. Hajibabaei. 2012. Next-generation sequencing technologies for environmental DNA research. Molecular Ecology 21:1794–1805.
Stein, E., B. P. White, R. D. Mazor, P. E. Miller, and E. M. Pilgrim. 2013. Evaluating ethanol-based sample preservation to facilitate use of DNA barcoding in routine freshwater biomonitoring programs using benthic macroinvertebrates. PLoS ONE 8:e51273.
Thomsen, P. F., and E. Willerslev. 2015. Environmental DNA — An emerging tool in conservation for monitoring past and present biodiversity. Biological Conservation 183:4–18.
Yahr, R., C. L. Schoch, and B. T. M. Dentinger. 2016. Scaling up discovery of hidden diversity in fungi: impacts of barcoding approaches. Philosophical Transactions of the Royal Society B: Biological Sciences 371:20150336.
Yu, D. W., Y. Ji, B. C. Emerson, X. Wang, C. Ye, C. Yang, and Z. Ding. 2012. Biodiversity soup: metabarcoding of arthropods for rapid biodiversity assessment and biomonitoring. Methods in Ecology and Evolution 3:613–623.
Zhou, X., Y. Li, S. Liu, Q. Yang, X. Su, L. Zhou, M. Tang, R. Fu, J. Li, and Q. Huang. 2013. Ultra-deep sequencing enables high-fidelity recovery of biodiversity for bulk arthropod samples without PCR amplification. GigaScience 2:4.
